The aim of structural biology is to provide a better understanding of biological systems for the advancement of medicine, pharmaceuticals etc. resulting in improved quality of life. One key to building this understanding is microscopy, of which X-ray microscopy is a technique that can be used to image structures down to the sub-micron level.
X-rays have the advantage over other microscopy techniques such as light and electrons in that they can image much thicker specimens on account of their relatively higher levels of permeability. This makes X-ray microscopy an excellent technique for seeing the inner structures. However, imaging biological materials is not always so easy, as many of the components have very similar densities. X-ray-based imaging technologies are strongly influenced by density and structures with similar densities provide low contrast, making imaging difficult.
Increasing Contrast between Neighbouring Structures
Strategies do exit to increase the contrast between neighbouring structures such as:
- Staining with contrast agents.
- Applying a median/Gaussian-based noise filter (denoise)
- Phase retrieval
In this article we will focus on the phase retrieval method.
The Rigaku nano3DX is a laboratory-based high-resolution X-ray microscope that uses a high-brilliance X-ray generator, a quasi-parallel beam technology and a high-resolution X-ray camera. The parallel beam geometry is advantageous over other systems in that it is immune to image blurring and loss of resolution caused by X-ray focus broadening, vibration and drift. The nano3DX also offers characteristic or continuous X-rays from four different target materials (Cr, Cu, Mo, W, although the Cu and Mo targets are sufficient for most biological specimens.
In the nano3DX measurement with a hard X-ray source, a projection image of weakly absorbing specimens shows characteristic refraction-derived fringes using phase retrieval techniques. In the energy region of hard X-rays, the contribution of the phase to the X-ray contrast is much larger than that of the absorption, indicating that the contrast may be efficiently improved if phase retrieval is successful.
Example 1 – Human Hair
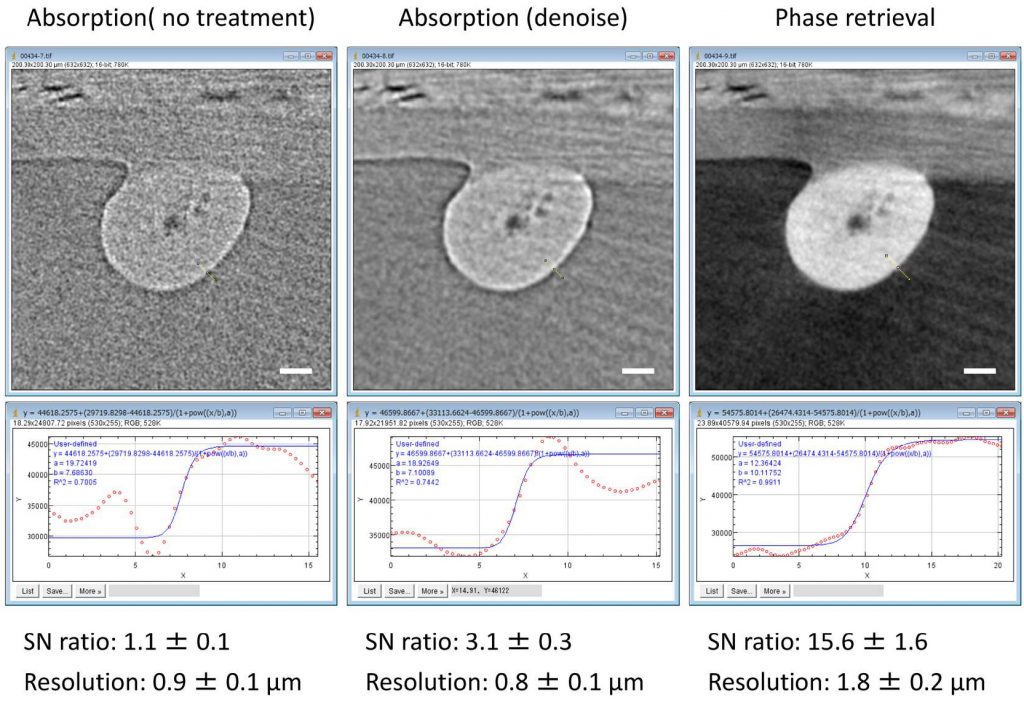
Figure 2 shows an example for contrast enhancement of an unstained human hair by the phase retrieval. In this case, the sample to detector distance was 4 mm and data was collected with a voxel size of 0.63 µm. The algorithm used for phase retrieval was the Paganin method assuming that the sample was made of organic materials (δ/β = 600). While the original CT slice shows fringes on the substance boundaries, the fringes disappear in the phase-retrieved image with concomitant enhancement of the signal to noise (SN) ratio by about ten-fold. As a result, the low-density medulla structures at the core of the hair sample are clearer in the phase-retrieved image. This type of contrast enhancement would then allow for software-assisted image processing, such as automated segmentation for regions of interest.
Example 2 – Pansy Seed
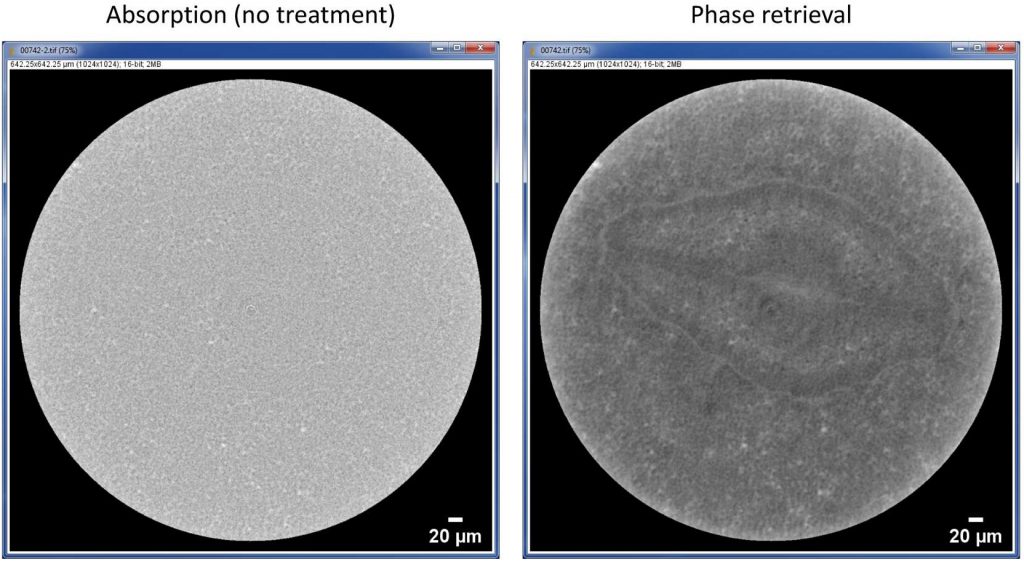
Another example is shown in Figure 3. This is a live observation of an unstained pansy seed fixed on the specimen stage to avoid any possible movement of the specimen. Imaging conditions are the same as for human hair example, except for a longer sample to detector distance of 7 mm. After contrast enhancement by phase retrieval, the structure of embryo and its inner structures are clearly observed, while almost no structure is seen in the original CT slice.
Summary
These examples illustrate how the phase retrieval method can be used with X-ray microscopes such as the Rigaku nano3DX to image biological materials down to the cellular level.
For more details please refer to “Application of the nano3DX X-ray Microscope to Biological Specimens”, Rigaku Journal Vol. 36 (1) pp. 33-36, 2020.

