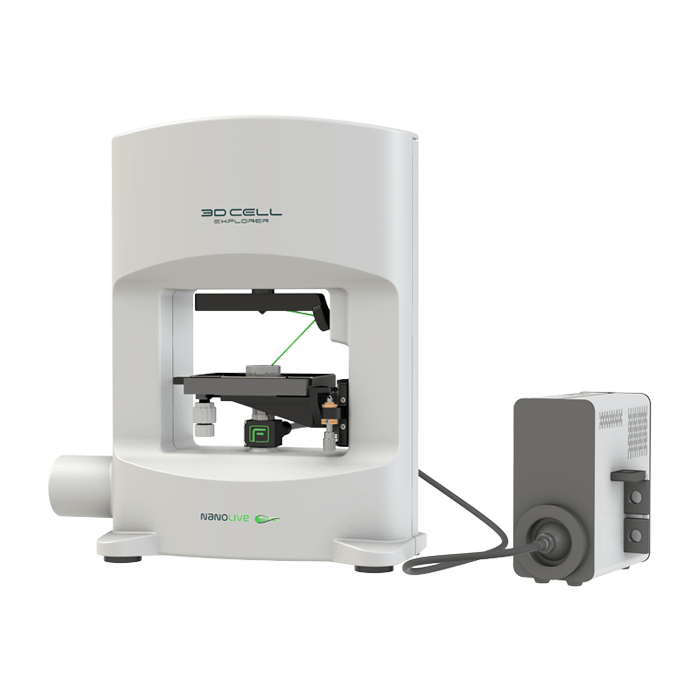
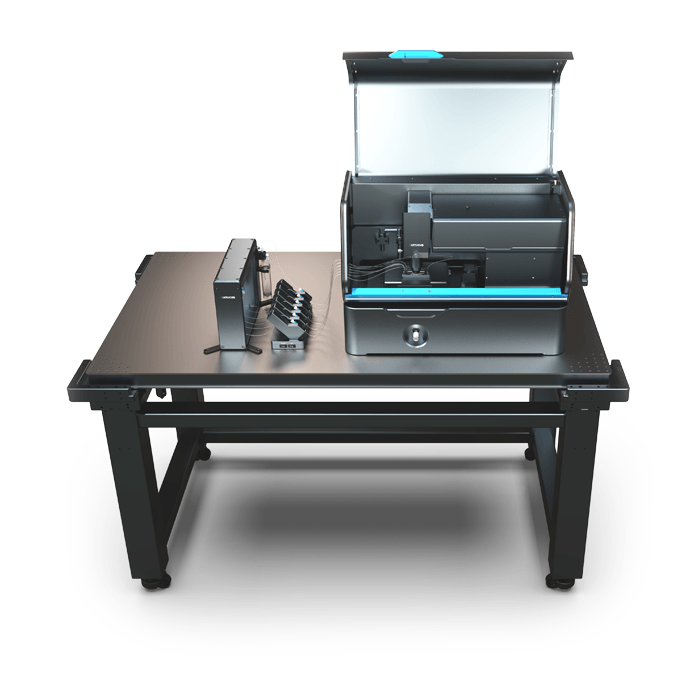
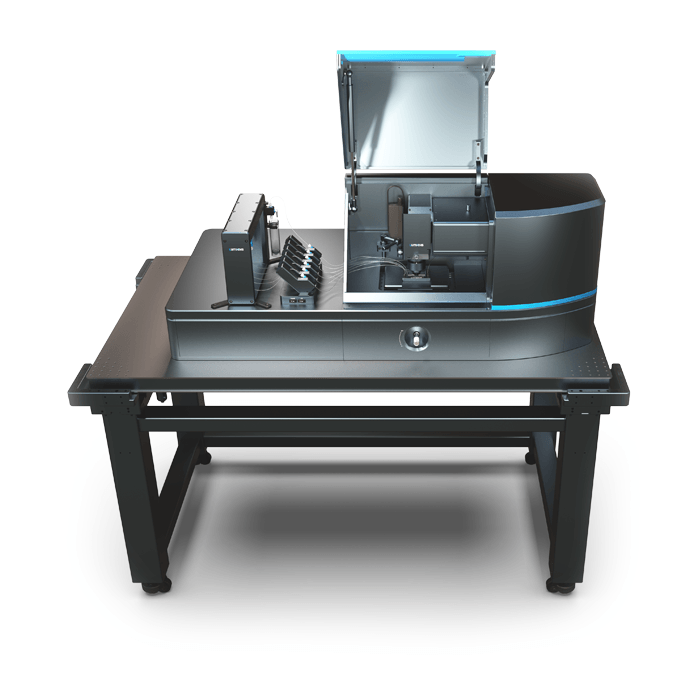
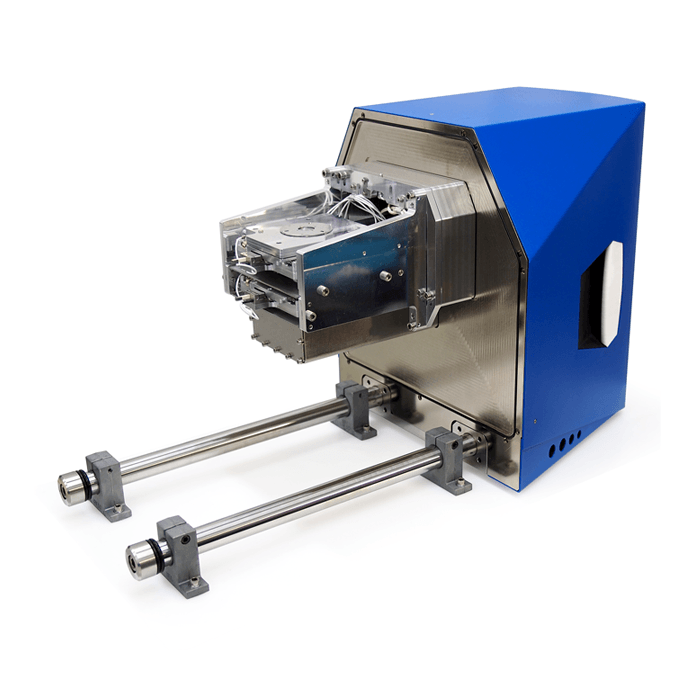
3D Cell Explorer – Live Cell Imaging Platform
Holotomography Microscopy

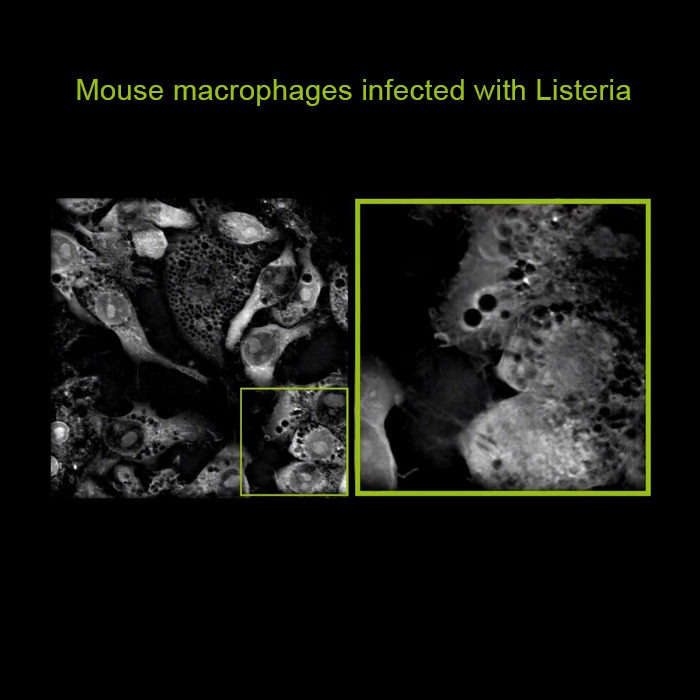
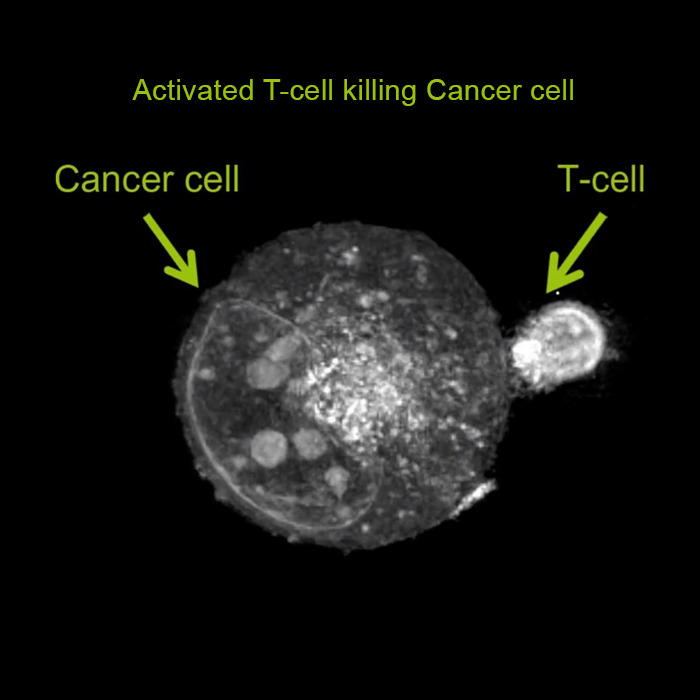
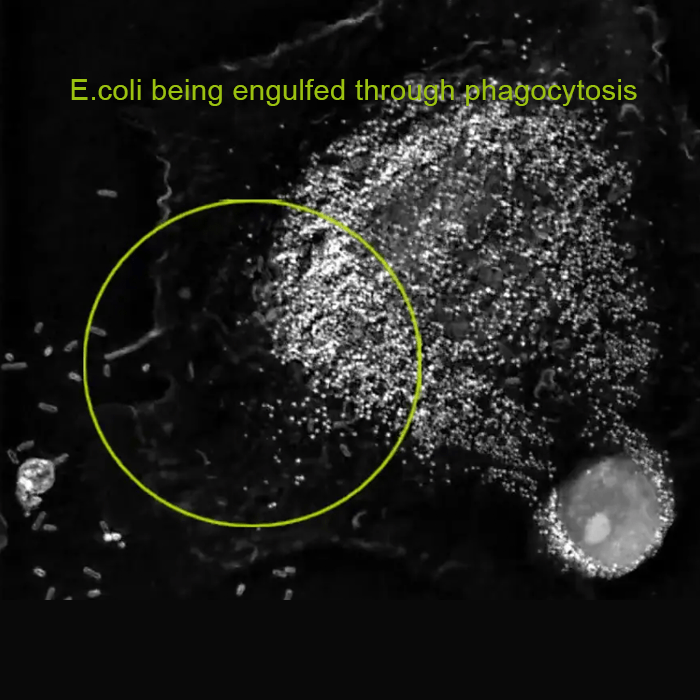
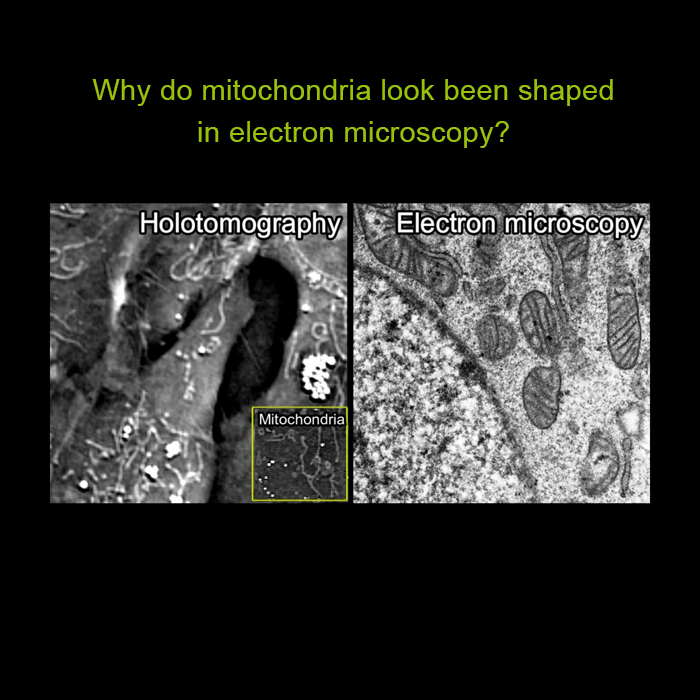
For the first time ever explore the inside of a living cell in 3D without the need for labeling. Take your Cell Research to the Next Dimension
-

Key Features
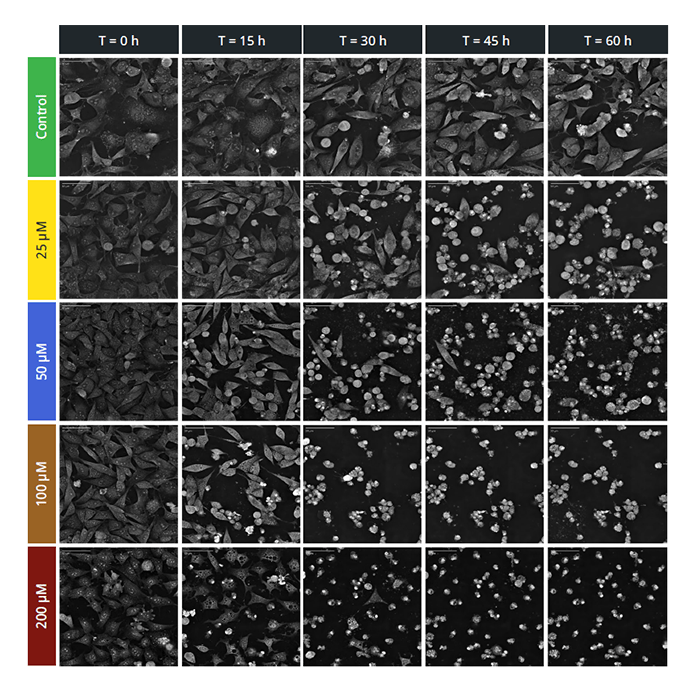
- Non- invasive 3D characterization - Live cell imaging in physiological conditions without any bleaching or phototoxicity
- Label-free 4D continuous Observation - Measurement of cell processes from seconds to weeks
- Multiplexing, Resolution to below 200nm in X-Y - High resolution and high sensitivity characterization of multiple cell organelles based on their refractive index.
- Full 3D scan in under 2 seconds - Fast imaging for capturing dynamic processes
- Long term live cell capable - Heat, humidity, CO2 and O2 control for long term stable timelapse imaging of even the most sensitive cells
Product Review
-
“It is absolutely fantastic! Its ease of operation, intuitive nature, compact size, rapid imaging and no need for stains make this a system I would certainly recommend and it would certainly support many different kinds of research. In fact, this system would be valuable for secondary students studying biology too.”
Wojtek Chrzanowski, MSc, PhD, DScSenior Lecturer, Faculty of Pharmacy and the Australian Institute for Nanoscale Science and Technology at the University of Sydney, Australia
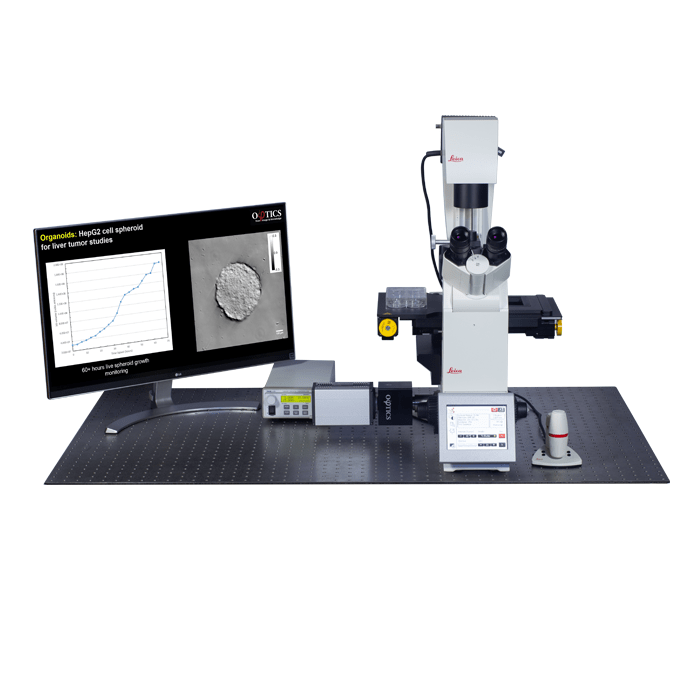
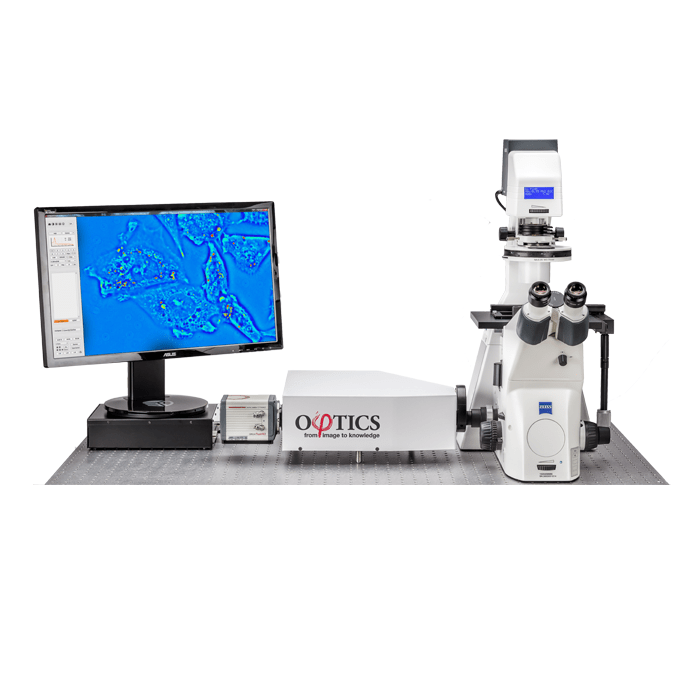
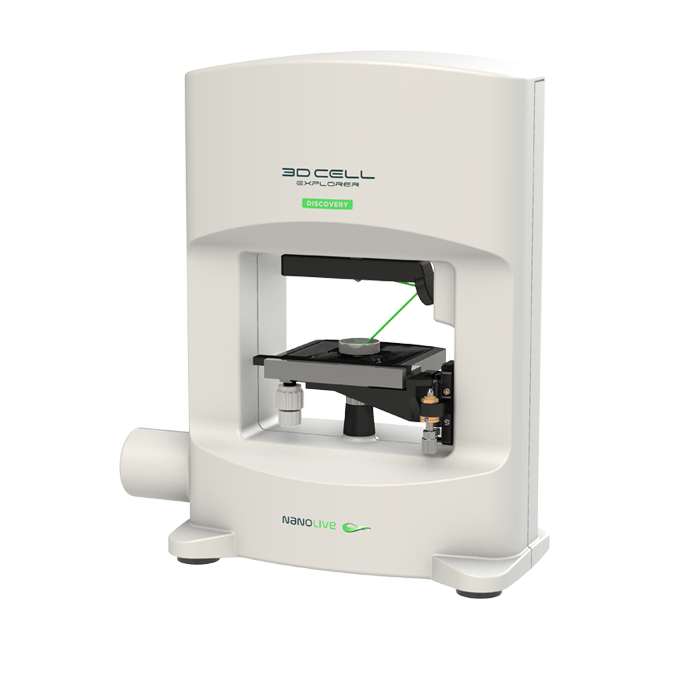
3D Cell Explorer
A disruptive technology which, for the first time ever, allows users to explore the inside of a living cell in 3D without the need for any labeling or other invasive methods.
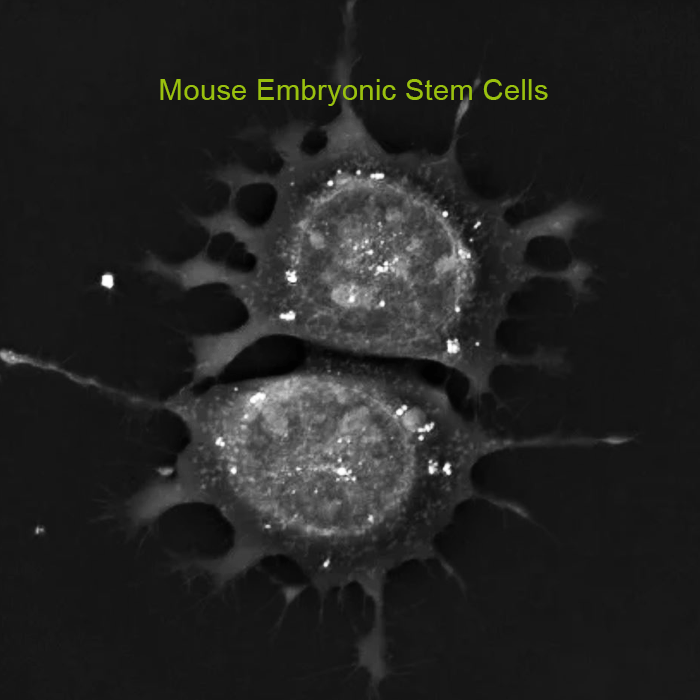
The 3D Cell Explorer is a high speed, high resolution and non-invasive tool that can look deep inside biological systems. This allows users to record stunning 3D images of entire cells in just seconds and with a higher resolution than any conventional light microscope on the market.
Non-invasive characterisation of live cells in physiological conditions: The 3D Cell Explorer measures the quantitative refractive index of cell organelles in seconds. Hence, it is possible to do non-invasive in vitro imaging of almost any kind of cells with up to 30 μm depth of reconstruction. This allows for biological features to be segmented based on their physical characteristics.
Label-free continuous observation of cell processes from seconds to weeks: Study cell life cycle processes of growth, division & death in 3D and 4D. Thanks to a dedicated top-stage incubator you can monitor cell compartments and their kinetics and dynamics in real-time at every second without interfering with their natural physiology.
Multiplexing: High resolution and high sensitivity characterization of multiple cell organelles based on their refractive index. Explore and measure up to 8 cell organelles simultaneously with unprecedented detail and resolution, marker-free and preparation-free based on their own physical density.
3D Data Sets: Multiplexing / unique organelle segmentation / Quantitative data analysis
Save Experimental Time: No sample preparation / short setup time / fast & easy acquisition
-
Learn how to use the 3D Cell Explorer
- Resolution Δx,y = 200nm; Δz = 400nm
- Field of View 85 x 85 x 30 μm
- Tomography frame rate 0.5 fps 3D image rate with full self-adjustement
- Microscope Objective Air with 60x magnification
- Illumination Source Class 1 laser low power
(λ=520 nm, sample exposure 0.2 mW/mm2) - Accessible Sample Stage 60 mm of free access to the sample stage for sample manipulation
- Dimensions 38 cm x 17 cm x 45 cm

- All
- Biological Microscopy
- Bioprinting
- Cell Culture
- Confocal Microscopy
- Digital Microscopy
- DNA/RNA
- Electron Microscopy
- Fluorescent Microscopy
- Light Microscopy
- Live Cell Imaging
- Microscopy
- Molecular Biology
- Multiphoton Microscopy
- Optical Tweezers
- Pre-clinical Imaging
- Protein
- Quantitative Phase Imaging
- Small Molecule
- Super Resolution Microscopy
- TEM
- Tomographic Microscopy